How do cells that migrate collectively maintain their shape while moving within confined tissues?
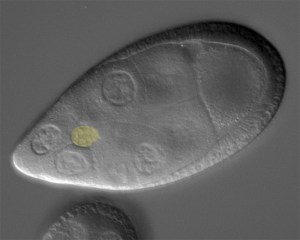
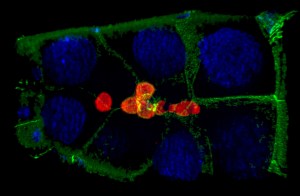
Cell collectives need to stay together and keep their shape, all while moving through confined environments. Border cells (light yellow in image) are 6-10 cells that move as a group between large “nurse cells” to navigate towards the oocyte (large cell to upper right in picture). We recently discovered a role for dynamic myosin at the outer edges of the border cell collective to resist the compression provided by the surrounding nurse cells. In unpublished work, we found that the shapes and positions of border cell nuclei are highly dynamic and reflect their position with the collective. Open questions include: What other proteins influence the shape of the collective? How do individual border cells maintain their shape during this process? How do individual cells within collectives deal with a relatively stiff nucleus when moving through such constricted spaces? And how does mechanical feedback between border cells and the nurse cells influence border cell migration?
Reading:
Dynamic myosin activation promotes collective morphology and migration by locally balancing oppositional forces from surrounding tissue. Aranjuez G, Burtscher A, Sawant K, Majumder P, McDonald JA. Mol. Biol. Cell 2016 27(12): 1898-1910. doi: 10.1091/mbc.E15-10-0744.
How do cells stay together and communicate while migrating as collectives?
How do normal and cancerous cells stay together to move as a group rather than as single cells? Our lab recently discovered that Protein Phosphatase 1 (PP1) activity inhibition causes border cells to round up, separate from each other and function as individual cells. We are currently investigating the mechanisms for how this genetic pathway controls adhesion between cells, maintains individual cell shapes, and keeps the border cell group moving. In addition, we are exploring the roles of the Rap1 small GTPase in regulating cell-cell adhesion within the collective that help border cells stay together and communicate while migrating. In an NSF Track 4 EPSCoR fellowship, we are building tools to manipulate Rap1 activity using light.
Reading:
K. Sawant§, Y. Chen§, N. Kotian§, K.M. Preuss, J.A. McDonald. Rap1 GTPase promotes coordinated collective cell migration in vivo.Mol Biol Cell. 2018 Nov 1;29(22):2656-2673. doi: 10.1091/mbc.E17-12-0752. Epub 2018 Aug 29. PMID: 30156466. §Equal contribution.
Y. Chen, N. Kotian, G. Aranjuez, L. Chen, C.L. Messer, A. Burtscher, K. Sawant, D. Ramel, X. Wang, J.A. McDonald. Protein Phosphatase 1 activity controls a balance between collective and single cell modes of migration. Preprint on BioRxiv. eLife 2020;9:e52979 DOI: 10.7554/eLife.52979
How do cells break away from epithelia to become migratory? And how do epithelia withstand growth of the tissue?
An unresolved but critical question is how groups of cells break away (detach or delaminate) from epithelia to begin their migratory journey. Moreover, epithelia need to be organized for proper function, especially for normal function of tissues and organs. Disruption of epithelia promotes tumor invasion and metastasis.
Border cells represent an ideal system to image this process in real-time and to manipulate the cells in their native environment. We recently discovered that the polarity protein and serine-threonine kinase Par-1 regulates two aspects of detachment: cell polarity-dependent remodeling of adhesion and myosin-dependent contraction. Currently we are investigating the role of a phosphatase complex in promoting cell fate for detachment and a small GTPase (Rap1) in promoting the correct number of cells to become migratory border cells. We also discovered that Rap1 is required for cell survival and organization of the epithelia during egg chamber growth.
Reading:
PAR-1 kinase regulates epithelial detachment and directional protrusion of migrating border cells. McDonald JA, Khodyakova A, Aranjuez G, Dudley C, Montell DJ. Curr Biol. 2008 Nov 11;18(21):1659-67. doi: 10.1016/j.cub.2008.09.041.
Par-1 controls myosin-II activity through myosin phosphatase to regulate border cell migration. Majumder P, Aranjuez G, Amick J, McDonald JA.Curr Biol. 2012 Mar 6;22(5):363-72. doi: 10.1016/j.cub.2012.01.037. Epub 2012 Feb 9.PMID: 22326025
Rap1 promotes epithelial integrity and cell viability in a growing tissue. Messer CL, McDonald JA.Dev Biol. 2023 Sep;501:1-19. doi: 10.1016/j.ydbio.2023.05.006. Epub 2023 Jun 2.PMID: 37269969
Interplay of polarity and cytoskeletal regulatory proteins in migration
How do proteins that control cell polarity interface with the cytoskeleton during cell migration? Do these proteins function differently in collective cell migration? To address these questions, we undertook an RNAi screen for genes required for collective border cell migration. We particularly targeted genes that encode PDZ- (PSD95/DLG/ZO-1) domain containing proteins, because these proteins serve as large protein scaffolds that modulate cell polarity, cell junctions and the cytoskeleton. New work in the lab is identifying genes, through cell-specific (and migration stage-specific) RNA-sequencing, whose transcripts are upregulated (and downregulated) during different phases of migration. We hope to identify networks of genes that control different aspects of collective cell migration, including polarity, cytoskeleton, and adhesion genes, among others.
Reading:
On the role of PDZ domain-encoding genes in Drosophila border cell migration. Aranjuez G, Kudlaty E, Longworth MS, McDonald JA. G3 (Bethesda). 2012 Nov;2(11):1379-91. doi: 10.1534/g3.112.004093.
Dynamics of cell polarity in tissue morphogenesis: a comparative view from Drosophila and Ciona. Veeman M.T. and McDonald J.A. F1000Research. 2016 Jun 2; 5. pii: F1000 Faculty Rev-1084. doi: 10.12688/f1000research.8011.1. eCollection 2016.
K. Sawant§, Y. Chen§, N. Kotian§, K.M. Preuss, J.A. McDonald. Rap1 GTPase promotes coordinated collective cell migration in vivo.Mol Biol Cell. 2018 Nov 1;29(22):2656-2673. doi: 10.1091/mbc.E17-12-0752. Epub 2018 Aug 29. PMID: 30156466. §Equal contribution.
Application of Drosophila border cell model to tumor invasion
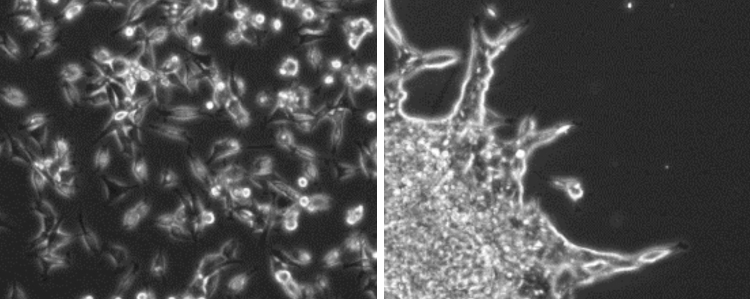
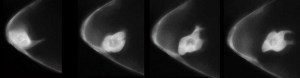
Using genes we have already discovered, along with new RNAi screens, we are applying our discoveries in border cell collective migration to human tumor invasion. This is a collaborative project with several basic and translational cancer researchers at the Cleveland Clinic Lerner Research Institute. We have found that some patient-derived tumor cell lines invade as single cells (left panel) in culture, whereas others invade in finger-like ‘collective’ strands (right panel). We are using genes found in border cells to identify those upregulated in human glioma patients. In one project, we identified conserved cell adhesion genes whose expression correlated with glioma patient survival and are required for border cell migration (Kotian et al., submitted). In new unpublished work, we are beginning to use Drosophila larval tumor models to test these same genes in collective tumor metastasis. Using these models, we hope to discover genes that control the transition between collective and single cell invasion and understand the impact these genes have on disease progression.
Reading:
Identifying conserved molecular targets required for cell migration of glioblastoma cancer stem cells. Volovetz J, Berezovsky AD, Alban T, Chen Y, Lauko A, Aranjuez GF, Burtscher A, Shibuya K, Silver DJ, Peterson J, Manor D, McDonald JA, Lathia JD. Cell Death Dis. 2020 Feb 26;11(2):152. doi: 10.1038/s41419-020-2342-2. PMID: 32102991
A Drosophila RNAi screen reveals conserved glioblastoma-related adhesion genes that regulate collective cell migration. Nirupama Kotian, Katie M. Troike, Kristen N. Curran, Justin D. Lathia, Jocelyn A. McDonald. 2021 bioRxiv Preprint. doi.org/10.1101/2021.08.09.455704. In press (2021): G3: Genes|Genomes|Genetics 2022 Jan 4;12(1):jkab356. https://doi.org/10.1093/g3journal/jkab356